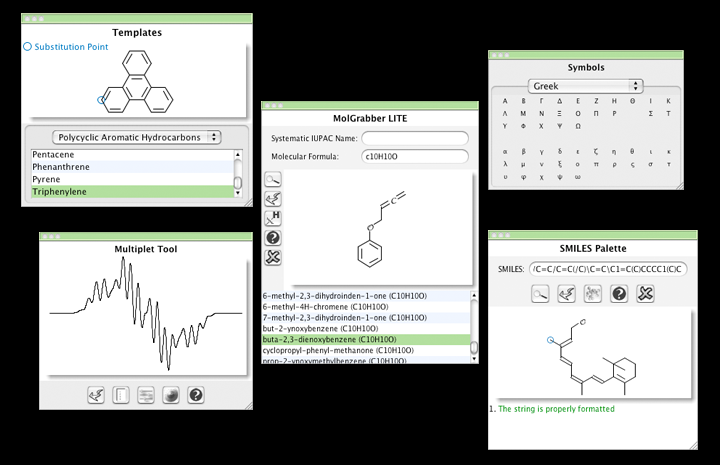
Capability to update Data Analysis graphs after having changed the order of the stacked spectra.After assigning a 2D peak, the correlated assignments are added to the applicable 1D peaks.New graphical assignment labels for 1D and 2D, which can be permanent as well as activated on hovering, to allow the user to more easily keep track, with a quick inspection, of peaks which have and have not been assigned.Īssignments are displayed directly on the chemical shift position.Improvements in the Assignment feature The assignment features in Mnova have been greatly enhanced with the objective to very significantly improve the workflow of the chemist when assigning multiple spectra sets. Copy the multiplet report from the table by using Ctrl+C.Capability to show number of multiplets as a floating point number in the multiplet label.You will be allowed to select the minimum area and what kind of peaks (Compound, Solvent, artifacts, etc) will be taken into account for the multiplet analysis. New Multiplet Analysis Options (independent of peaks and integrals settings).Here you can see an example of a triplet which is hidden under the solvent signal:.Multiplet AnalysisOnce again, Multiplet Analysis benefit directly from the exploitation of GSD capabilities to carry out the automatic analysis, with the enhanced peak picking capabilities resulting in much more reliable automatic multiplicity identification and labeling. This is very useful if you need for example to determine relaxation times from a series of 1D spectra, to be able to compare the integral values between the different traces. It will be possible to apply the same integral normalization to all the spectra (in the current document) or to all the traces of a stack plot by selecting the applicable option, as you can see below. Capability to compare integral values of different spectra.In the example below we have excluded the solvent signal for the integration. This will give the user the ability to select what kind of peaks will be taken into account for the integration.Integration The integration algorithm in Mnova also benefits from the exploitation of GSD capabilities, with GSD generated integral values becoming the new default in Mnova. NMR Peaks & Snap also pastes annotations.A series of graphical display enhancements relating to peaks are now implemented in Mnova, including:Ĭapability to change the color peak curves (depending on the type and flags)Ĭapability to change the color peak labels (or use the same curve colors).Peak labels are also hot areas which can be selected and interacted with just by right clicking on it (NMR peak context menu).įrom here you can edit the Peak Type, use the curve color for the peak label, run a Peak Search in the DB, etc.The user now enjoys the capability of selecting which kinds of peaks are to be shown/hidden (Compound, Solvent, artifacts, etc). 
For example, hovering the mouse over the peak label (or on any table) will highlight the peak in the spectrum and allow the user to interact directly with that peak or automatically scroll up/down the table to the position of currently highlighted peak in the spectrum just by clicking on it Peak curves are now active objects which can be interacted with, and three-way highlighting of peaks table, peak labels and the peak themselves has been implemented.The default automatic Peak Picking algorithm is now based on GSD, rendering enhanced resolution, identification of overlapped peaks and also including autodetection of solvent and impurities.Įxample of the enhanced resolution achieved by the new Peak picking algorithm, now capable of identifying shoulders on large peaks as smaller peaks and of labelling and include them in multiplet analysis. This is a major change to Mnova NMR functionality.New powerful and more accurate algorithms for Peak Picking, Integration and Multiplet Analysis A new refactoring for the Peak Picking, Integration and Multiplet algorithms has been implemented in Mnova 7 with the objective to get more accurate, detailed analysis and to minimize the need for user interaction.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
